Previously on Hapax Legomenon…
I ordered a test from 23andMe.com, spit in a tube and sent it back. Six weeks later, I have results in my browser. I had made some educated guesses about what I might find. We’ll see how I did.
As a male, I got to have both my Y-DNA and my mtDNA tested. Y-DNA traces my father’s line back through his father and so on. mtDNA does the same through my mother and her mother, etc. What you get is a line connecting you to the human male and female most recent common ancestors (MRCA). Certainly not a complete picture, but an illuminating one nonetheless.
I had predicted that my mtDNA would fall in haplogroup M, haplogroup R, or haplogroup U, but I couldn’t be more specific than that. I had predicted that my Y-DNA would fall in haplogroup R, probably subclade R1a1.
So here we go.
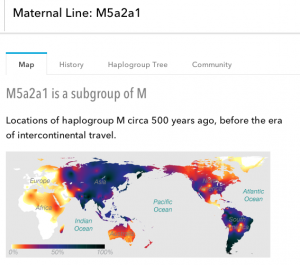
On the mtDNA front, I was right–sort of. As right as I could be when I hedged with three choices. My maternal line falls in haplogroup M5a2a1, one of the many subclades of the big group M. M is a huge lineage. It was one of the children of L3, the line that came out of Africa, and has been in India for 50,000 years! My maternal ancestors were among the first modern humans to make it to the subcontinent and have been there ever since.
M5 is one of the most diverse subclades of M. I haven’t been able to find much info on M5a2a1 specifically, but the 23andme blurb about M says that there is an especially diverse corridor of M lineages in northeastern India and Bangladesh. The state of Odisha is a little ways south of that, but M5a is apparently found in high concentrations of the Gadaba people of that state. That might mean there’s some tribal ancestry way back, and maybe there’s something to my grandmother’s story after all. Whatever the case, it seems that I have an extremely old line going back to that part of the world.
On the paternal front…
Yes. I am a droid.
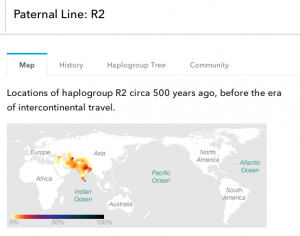
So I was actually wrong about my paternal DNA, though I was almost right. I’m actually in haplogroup R2 instead of R1a1. I had to spend some time dissecting what this meant.
R2 is found in many of the same Indian populations as R1 lineages, just in lower concentrations: R2 is much rarer (aren’t I just a special little snowflake?). While their R1 brothers went off and proliferated around Europe, R2 lineages stayed closer to their South/Central Asian origins, and R2 is common in India, Iran, and the Caucusus Mountains.
It’s definitely one of the later migrations into the Indian Subcontinent and possibly entered the area around the same time as the R1 lineages. Unfortunately, there seems to be a lot less data out there on R2 than there is on R1a or R1b.
From doing some research, I found out that what 23andMe is calling R2 is more properly termed R2a, and is due to a specific marker M-124. Much of the history of R2a mirrors that of R1a. They have similar distributions in similar Indian populations, and both may have originated in either South Asia or Central Asia and may have moved back and forth through history. I’ve read some talk that the spread of R1a may be correlated with the spread of Indo-European languages into the subcontinent. In that case, those migrating populations may have had some R2 in them as well, but there was probably also R1a and R2a present in the Indian Subcontinent before them.
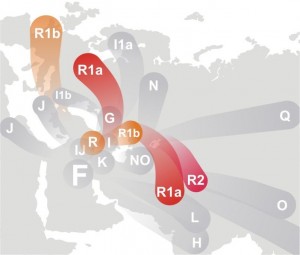
Y-Chromosome haplogroup diffusion: from Wikipedia
Overall, it seems that I have two very distinct lines in me: one that entered India very early and stuck around in the jungles and on the coast, and another that, 40000 years later, followed the typical migration/invasion route and rolled in from the northwest (and then rolled out and maybe in again). Everything comes over the Khyber Pass.

There were probably fewer trucks when my Y-chromosome entered the area.
I share bits of DNA with a few other 23andMe users, mostly Indians, but also one Romanian and one Brit (not a South Asian immigrant). I don’t know if that means I’m whiter than I thought or they’re more Asian than they thought.
According to their standard estimate, 23andMe says that I am 99.2% South Asian and 0.1% East Asian.
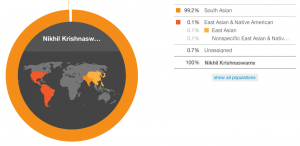
My ancestry composition (standard estimate). Ignore the red all over the Americas–they group East Asian and Native American together but the East Asian’s what’s relevant here.
Unfortunately, they don’t break these categories down into nationalities the way they do with Europe, so I can’t get more specific. According to their survey stats, about 87% of their users identify as European, which doesn’t make it easy for brown people like me. I need to get more South Asians to spit in a tube.
According to their conservative estimate, the South Asian percentage drops slightly and according to their speculative estimate, a tiny bit of European creeps in there.
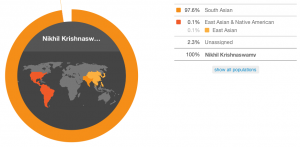
My conservative estimate.
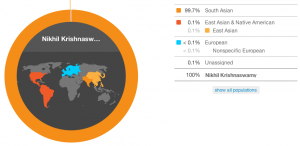
My speculative estimate. This chart is colorful and pretty, so I think I like it the best.
When I view my genome, the East Asian chunk is contiguous, which I would think suggests it all comes from one person. Assuming this and that that chunk isn’t just lucky to have survived dozens of generations unchopped, that person would have been my great-great-great-great-great-great-great-great-grandparent (that is, my great-grandparent’s great-grandparent’s great-grandparent’s parent, or ten generations before the present). And if you go back some more generations, you may find a European or two. The possibly European bits of my DNA are dotted all over my genome, suggesting multiple originators further back in time. So assuming an average generational length of 25 years (less going maternally, more going paternally), around the early 18th century is the latest time where you might find anything in my ancestry that would be considered outside the Indian subcontinent.
Clearly, I’m mostly South Asian (no surprises there), but with indicators that point both west and east, which I think is pretty damn cool, though I wish there were ways to know more.
I’m also 2.7% Neanderthal, which is slightly high for a South Asian (at least, those South Asians who are 23andme members, which is a small sample size). This might also be indicative of some West Eurasian ancestry in there somewhere.
I’d like to think that somewhere back in my ancestry there was some badass Central Asian horseman and a charming Indus Valley priestess who seduced him with the wonders of indoor plumbing or something, but, given that this is ancient history, it probably turned out more violent and creepy than I ever care to consider.
Overall, I learned some interesting things–most notably about my paternal great-grandfather’s origins, whose line I knew the least about going into this. 23andMe provides some interesting data, and some speculations which should probably be taken with a grain of salt. Their sample size for some things is small, heavily biased toward Europeans (which is a different issue), and based in part on self-reported data. But in the end, I had a lot of fun with this.
My fiancée also got her DNA tested. She appears to be mostly British, Dutch, Scandinavian, and Ashkenazi. Ashkenazi is grouped with Europe so unfortunately obscures any potential Middle Eastern connections of this Jewish group. Her maternal line was a lot more on the move than mine. Her mtDNA haplogroup is H1g, a very rare variant of the common European H1 that arose in the Iberian peninsula about 13,000 years ago and followed the retreating glaciers north. H1g is common among Scandinavians, but also Lebanese Druze, and indigenous groups in Japan (possibly Ainu?). The 23andMe calculator, though, seems pretty certain that she’s mostly is old European.
One last thing: I got no famous people in my haplogroups! Y-DNA R1s get everyone from Tsar Nicholas II to Stephen Colbert but R2s get nobody. And for all the range that mtDNA M got, I don’t see anyone famous who’s an M5. Not even an asshole! Guess I have to become famous now and fix that–at least famous enough to warrant a Wikipedia page.

Pingback: DNA Test Reveals White Girl Is White | Hapax Legomenon
This was an interesting read. I am a Tamil Iyengar (vadagalai) as well and my husband is white. He had his dna done with 23andme and I am thinking of getting mine done. We are really curious to have our son and daughter sampled as well!